1. Rcpp sugar
In a previous chapter we explained how Rcpp allows users to interface R with C++ in a convenient way. Here we cover Rcpp sugar, which consists of a set of functions and operators which make the C++ code written using Rcpp behave similarly to R code. In other words, Rcpp sugar allows us to write C++ code which looks similar to its R equivalent, but often more efficient.
This document follows closely the official Rcpp sugar vignette.
Vectorized operators and sugar versions of basic R functions
Consider the following Rcpp code for summing two vectors:
library(Rcpp)
sourceCpp(code = '
#include <Rcpp.h>
using namespace Rcpp;
// [[Rcpp::export(name = "vsum")]]
NumericVector vsum_I(const NumericVector x1, const NumericVector x2)
{
int ni = x1.size();
NumericVector out(ni);
for(int ii = 0; ii < ni; ii++){
out[ii] = x1[ii] + x2[ii];
}
return out;
}')First of all, let’s check whether it works:
d <- 1e5
x1 <- rnorm(d)
x2 <- rnorm(d)
y <- vsum(x1, x2)
max( abs(y - (x1+x2)) )## [1] 0It seems so. The Rcpp code above is quite lengthy: do we really need to write a for loop to simply add two vectors together? Thanks to Rcpp sugar, we don’t have to. In particular, Rcpp sugar uses operator overloading to vectorize many operations involving vectors represented using the Rcpp vector classes (NumericVector, IntegerVector, …). Hence, we can write vector addition in a much more compact form:
sourceCpp(code = '
#include <Rcpp.h>
using namespace Rcpp;
// [[Rcpp::export(name = "vsumVett")]]
NumericVector vsum_I(const NumericVector x1, const NumericVector x2)
{
return x1 + x2;
}')Let’s see whether this works:
y <- vsumVett(x1, x2)
max( abs(y - (x1+x2)) )## [1] 0It does! All four basic arithmetic operations +, -, * and / are vectorized, thanks to Rcpp sugar. Vector-scalar operations are also vectorized, for instance we can do \(x1 * 3.0 + 1.0\), which gives the same result as in R.
However, note that writing such simple arithmetic expression in Rcpp will not give you a big performance gain in most cases, for instance:
library(microbenchmark)
microbenchmark(R = x1 + x2, Rcpp = vsum(x1, x2), RcppSugar = vsumVett(x1, x2))## Unit: microseconds
## expr min lq mean median uq max neval
## R 127.5 426.75 521.981 431.75 440.45 3932.0 100
## Rcpp 159.5 513.10 636.968 519.30 528.20 13796.9 100
## RcppSugar 129.6 431.75 482.644 436.80 446.30 5428.8 100This is because basic arithmetic operations in R are computed using internal code, which is written in C or C++. Calling C++ from R has also some overheads, so the C++ code can actually become much slower than the R version, for instance:
d <- 1e2
x1 <- rnorm(d)
x2 <- rnorm(d)
microbenchmark(R = x1 + x2, Rcpp = vsum(x1, x2), RcppSugar = vsumVett(x1, x2))## Unit: nanoseconds
## expr min lq mean median uq max neval
## R 100 300 427 400 400 4000 100
## Rcpp 2300 2600 2879 2800 3000 6100 100
## RcppSugar 2200 2500 2840 2700 2900 12100 100Here Rcpp is much slower than R, because the computational effort is limited (the vectors x1 and x2 are short) and it is dominated by the cost of calling C++ from R. Closing this brief digression on the dangers of using Rcpp without a clear idea of where the performance gains will come from, we go back to Rcpp sugar.
Rcpp sugar provides vectorized versions of the main logical operators >, <, >=, <=, ==, &, | and !. For instance, we can define the function:
sourceCpp(code = '
#include <Rcpp.h>
using namespace Rcpp;
// [[Rcpp::export(name = "inRange")]]
LogicalVector inRange_I(const NumericVector x, const NumericVector u, const NumericVector l)
{
return x > l & x < u;
}')which checks which elements of x falls in the range defined by l and u. Example usage:
set.seed(5151)
n <- 5
u <- rep(0.8, n)
l <- rep(0.2, n)
x <- runif(n)
data.frame(x = x, inside = inRange(x, u, l))## x inside
## 1 0.78699386 TRUE
## 2 0.07425964 FALSE
## 3 0.93492781 FALSE
## 4 0.22155905 TRUE
## 5 0.47941927 TRUEAs in base R, some operators can be used also in a “unary” sense, for example !x will negate a LogicalVector and -x will change the sign of a NumericVector or IntegerVector.
Rcpp sugar also provides Rcpp versions of many basic R functions, including:
- logical operations such as
ifelse,all,any, …; - arithmetic operation such as
sign,sqrt,diff,cumsum, …; - special functions such as
gamma,beta,choose, …; - stat summaries such as
mean,median,range, …; - matrix operations such as
colSums,diag, …; - statistical distribution such as
dnorm,pgamma, …; - …
See here for a full list. This makes working with Rcpp very easy, if you are already familiar with R. For example, we can do:
cppFunction('
List exampleSugar(NumericVector x) {
NumericVector x1 = cumsum(x);
LogicalVector x2 = any(is_na(x));
NumericMatrix x3 = cbind(x1, x1);
List out = List::create(x1, x2, x3);
return out;
}')
exampleSugar( c(1, 0, -2, NA) )## [[1]]
## [1] 1 1 -1 NA
##
## [[2]]
## [1] TRUE
##
## [[3]]
## [,1] [,2]
## [1,] 1 1
## [2,] 1 1
## [3,] -1 -1
## [4,] NA NAwhich gives us the same results we would get in base R.
Random number generation
Rcpp sugar provides C++ level access to the random number generator (RNG) available in R. For example, we can do:
cppFunction('
NumericVector rbeta_rcpp(const int n, const int a, const int b) {
return rbeta(n, a, b);
}')
hist(rbeta_rcpp(1e3, 1, 3))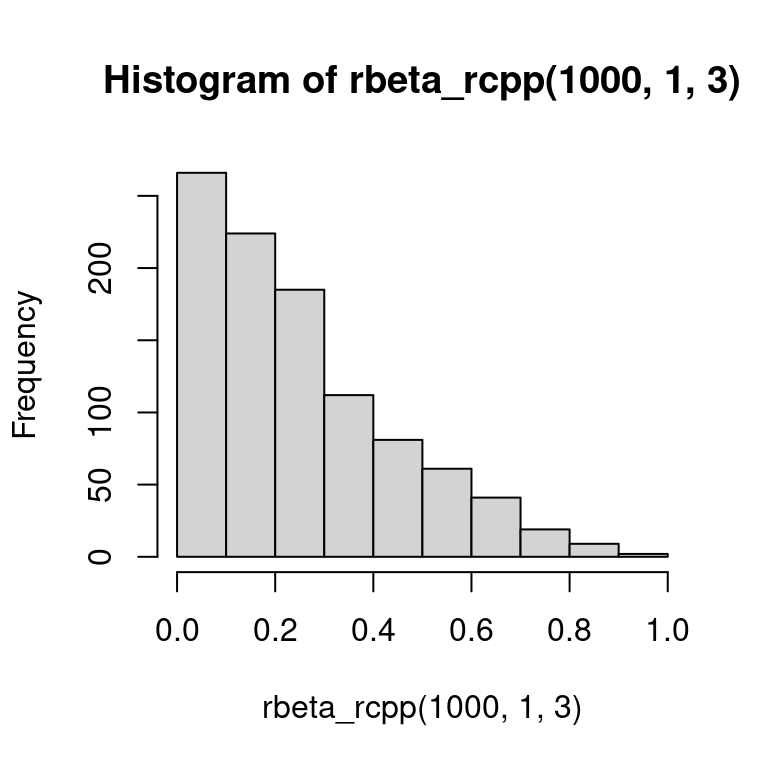 Given that
Given that Rcpp is using R’s RNG, setting the seed within the R session will make the results reproducible and identical to what we would get with base R:
set.seed(321)
rbeta_rcpp(5, 1, 3)## [1] 0.5159567 0.3422122 0.2875003 0.2889146 0.1782904set.seed(321)
rbeta(5, 1, 3)## [1] 0.5159567 0.3422122 0.2875003 0.2889146 0.1782904cppFunction adopts the Rcpp namespace by default hence, within rbeta_rcpp, the line rbeta(n, a, b) is equivalent to Rcpp::rbeta(n, a, b). Rcpp::rbeta outputs a NumericVector of length n. If, instead, we want to simulate random variables one at the time, we can use functions defined in the R namespace. For example:
cppFunction('
NumericVector rbeta_R(const int n, const int a, const int b) {
NumericVector out(n);
for(int ii = 0; ii < n; ii++) out[ii] = R::rbeta(a, b);
return out;
}')
set.seed(321)
rbeta_R(5, 1, 3)## [1] 0.5159567 0.3422122 0.2875003 0.2889146 0.1782904Note that R::rbeta returns a double, not a NumericVector. See here for a complete list of the statistical distribution available via Rcpp sugar.
As explained here and in the Writing R Extensions documentation, to use R’s internal RNG in C/C++ we need to call GetRNGState before using the RNG and PutRNGState before returning to R. The first call reads the state of the RNG (see ?.Random.seed), the second updates it. When we use R’s RNG within functions exported via the // [[Rcpp::export]] attribute we don’t have to call GetRNGState or PutRNGState, because Rcpp does it for us in the automatically generated C++ wrapper function. The // [[Rcpp::export]] attribute is used automatically by cppFunction and, in fact, if we recompile a previous example:
cppFunction('
NumericVector rbeta_rcpp(const int n, const int a, const int b) {
return rbeta(n, a, b);
}', rebuild = TRUE, verbose = TRUE)##
## Generated code for function definition:
## --------------------------------------------------------
##
## #include <Rcpp.h>
##
## using namespace Rcpp;
##
## // [[Rcpp::export]]
##
## NumericVector rbeta_rcpp(const int n, const int a, const int b) {
## return rbeta(n, a, b);
## }
##
## Generated extern "C" functions
## --------------------------------------------------------
##
##
## #include <Rcpp.h>
## #ifdef RCPP_USE_GLOBAL_ROSTREAM
## Rcpp::Rostream<true>& Rcpp::Rcout = Rcpp::Rcpp_cout_get();
## Rcpp::Rostream<false>& Rcpp::Rcerr = Rcpp::Rcpp_cerr_get();
## #endif
##
## // rbeta_rcpp
## NumericVector rbeta_rcpp(const int n, const int a, const int b);
## RcppExport SEXP sourceCpp_9_rbeta_rcpp(SEXP nSEXP, SEXP aSEXP, SEXP bSEXP) {
## BEGIN_RCPP
## Rcpp::RObject rcpp_result_gen;
## Rcpp::RNGScope rcpp_rngScope_gen;
## Rcpp::traits::input_parameter< const int >::type n(nSEXP);
## Rcpp::traits::input_parameter< const int >::type a(aSEXP);
## Rcpp::traits::input_parameter< const int >::type b(bSEXP);
## rcpp_result_gen = Rcpp::wrap(rbeta_rcpp(n, a, b));
## return rcpp_result_gen;
## END_RCPP
## }
##
## Generated R functions
## -------------------------------------------------------
##
## `.sourceCpp_9_DLLInfo` <- dyn.load('/tmp/Rtmp358fok/sourceCpp-x86_64-pc-linux-gnu-1.0.10/sourcecpp_b56220a9763/sourceCpp_13.so')
##
## rbeta_rcpp <- Rcpp:::sourceCppFunction(function(n, a, b) {}, FALSE, `.sourceCpp_9_DLLInfo`, 'sourceCpp_9_rbeta_rcpp')
##
## rm(`.sourceCpp_9_DLLInfo`)
##
## Building shared library
## --------------------------------------------------------
##
## DIR: /tmp/Rtmp358fok/sourceCpp-x86_64-pc-linux-gnu-1.0.10/sourcecpp_b56220a9763
##
## /opt/R/4.2.2/lib/R/bin/R CMD SHLIB --preclean -o 'sourceCpp_13.so' 'fileb5623898f60b.cpp'we see that the C++ wrapper contains the line Rcpp::RNGScope rcpp_rngScope_gen; which internally calls GetRNGState. PutRNGState will be called automatically when exiting the C++ wrapper function.
One case when we need to worry about the reading and writing the R’s RNG state, is when we want to do things manually, that is without using the // [[Rcpp::export]] attribute. To provide an example, define the function:
#include <Rcpp.h>
using namespace Rcpp;
RcppExport SEXP rnorm_manual(SEXP n) {
NumericVector out( as<int>(n) );
RNGScope rngScope;
out = rnorm(as<int>(n), 0.0, 1.0);
return out;
}in the “rnorm_manual.cpp” file, load it and test it:
system("export PKG_CXXFLAGS=`Rscript -e \"Rcpp:::CxxFlags()\"` &&
R CMD SHLIB rnorm_manual.cpp")
dyn.load("rnorm_manual.so")
set.seed(31)
.Call("rnorm_manual", 3)## [1] 0.05557024 -0.18423859 1.59576183set.seed(31)
rnorm(3)## [1] 0.05557024 -0.18423859 1.59576183Here we needed to create an RNGScope object, before using the RNG via rnorm. IMPORTANTLY the RNGScope must be declared AFTER declaring any of the outputs of the function (the vector out in this case). Failing to do so, for example declaring an output after the RNGScope object, will lead to very annoying intermittent crashes. Hence, it is generally preferable to use the Rcpp::export attribute, which deals with the RNG state automatically.